About DeepSpaceDB
DeepSpaceDB is an interactive database of spatial transcriptomics data. While existing spatial omics databases broadly collect large numbers of samples from many different platforms, DeepSpaceDB instead focuses on enabling a deeper analysis of data generated using the most popular platforms. The goals of DeepSpaceDB include:
- Enable both wet and dry researchers to easily explore and visualize publicly available spatial transcriptomics samples.
- Facilitate the generation of new testable hypotheses.
- Make possible the re-analysis of existing spatial transcriptomics data by providing downloadable data in consistently processed formats.
The various cell subpopulations that make up biological systems and organs have physiological mechanisms and operations that are closely linked to their spatial distributions and cellular interactions. Although single-cell RNA sequencing (scRNA-seq) continues to show progress, a critical practical challenge remains to release viable cells from entire tissues without causing cellular stress, apoptosis, or aggregation. However, the spatial resolution of gene expression information derived from intact tissue slices in their original physiological environment is made possible by spatial transcriptomics. It is possible to gain a variety of biological insights into tissue architecture and clarify how cells interact with their surroundings by analyzing spatial transcriptomics data. DeepSpaceDB is a database that provides comprehensive analysis of spatial transcriptomics data for human and mice using spatial data from various organs to understand the complex pathophysiological mechanisms. Our platform aims to provide advanced tools to enable researchers, users, and scientists to investigate gene expression patterns in relation to the spatial context of tissue architecture.
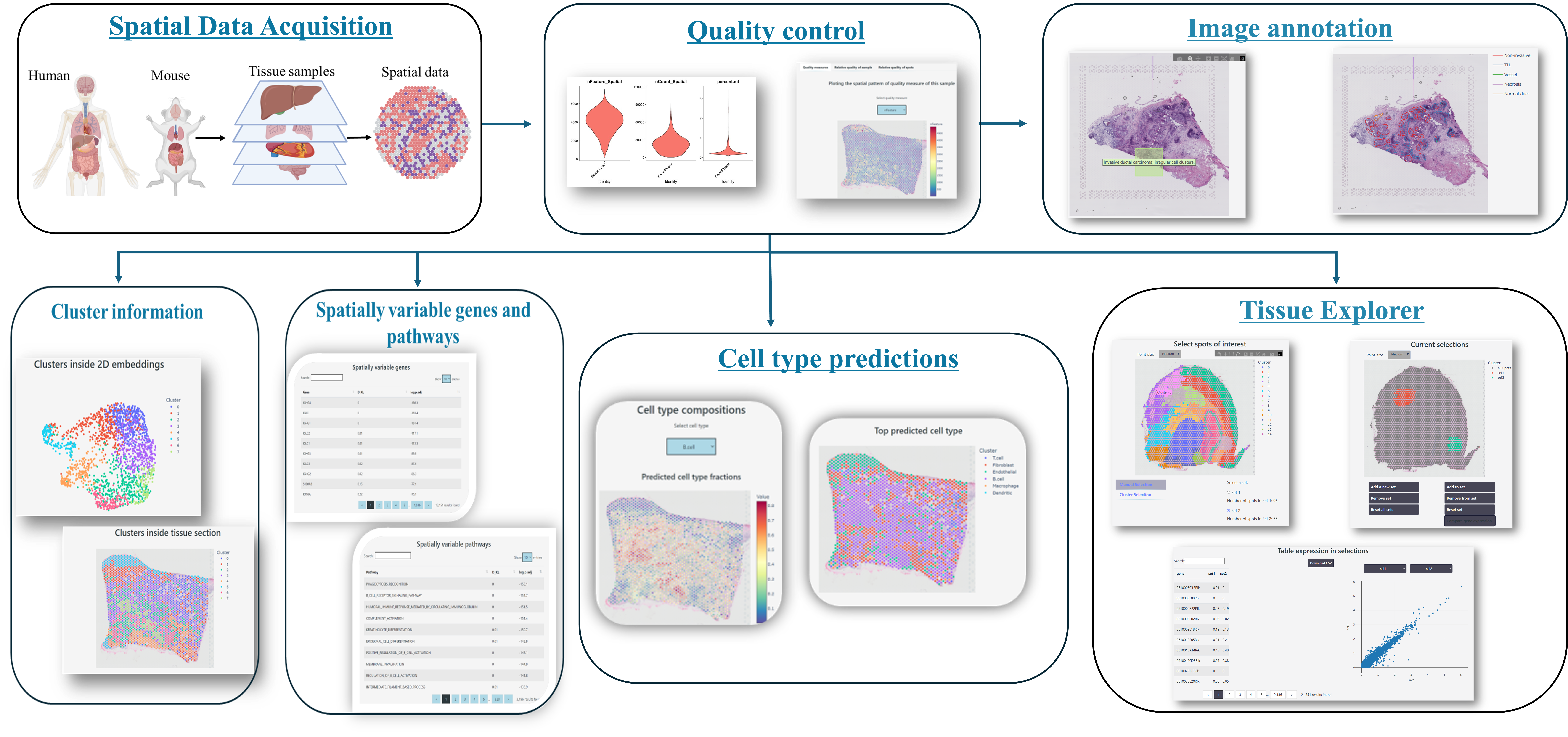
This database's key features consist of various detailed analysis that includes, quality control, identification of spatially variable genes, pathways enrichment, cell-type predictions, identifying similar spots across database. The robust data processing, quality assurance, and integration of various datasets form a fundamental component of our spatial transcriptomics database. Our goal is to make complex spatial transcriptomics analysis accessible to all learners through a user-friendly layout. Our database is made to easily and precisely support scientific study, regardless of the user's purpose, for example, examining cellular heterogeneity, discovering novel biomarkers, or investigating tissue organization.
Funding
This work is supported by JST NBDC Grant Number JPMJND2303 (Japanese - English) and by the Future Development Research Funding Program (FDRFP) of the Kyoto University Research Coordination Alliance.
Citation
Honcharuk V., Zainab A., Horimoto Y., Takemoto K., Diez D., Kawaoka S. and Vandenbon A., "DeepSpaceDB: a spatial transcriptomics atlas for interactive in-depth analysis of tissues and tissue microenvironments", Nucleic Acids Res., 2025, doi: 10.1093/nar/gkaf111